Predicting and hyperparameters tuning
Support Vector Machines (SVMs)¶
Introduction
- Support vector machines are models that learn to differentiate between data in two categories based on past examples
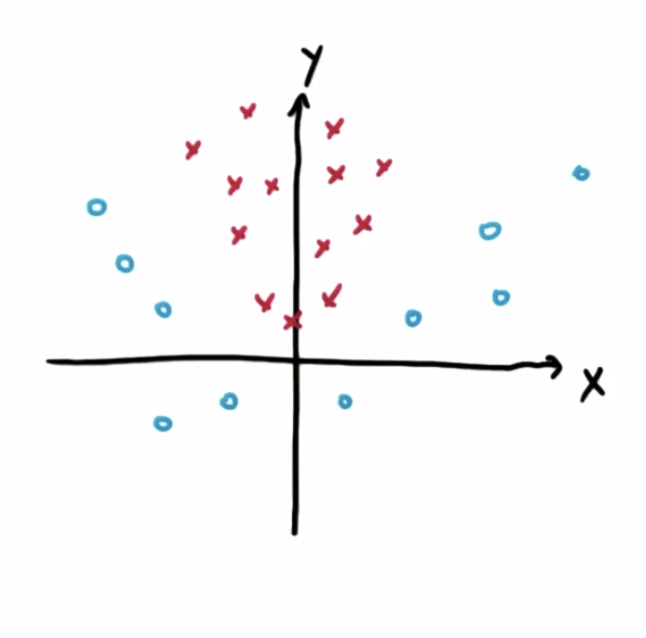
- We want to have the maximum margin from the line to the points as shown in the diagram and that is the essence of SVMs
- The points close to the decision boundary matters, the rest are not important
- Note that SVMs aim to classify correctly before maximizing the margin

- You can also project your data into a higher dimensionality and split them with a hyperplane
- Optimization problem for finding maximum margins uses quadratic programming
- We use a kernel to identify similarity amongst points
- All kernels must satisfy Mercer conditions
SVM using Scikit-Learn
In [1]:
from sklearn.svm import SVC
from sklearn.datasets import load_iris
from sklearn.cross_validation import train_test_split
from sklearn.metrics import accuracy_score
import seaborn as sns
In [2]:
# Create object
iris = load_iris()
# Create data
X = iris.data
y = iris.target
# Split data
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=0)
In [3]:
# Same 3 steps
# 1. Instantiate
# Default kernel='rbf'
# We can change to others
svm = SVC(kernel='linear')
# 2. Fit
svm.fit(X_train, y_train)
# 3. Predict
y_pred = svm.predict(X_test)
In [5]:
# Accuracy calculation
acc = accuracy_score(y_pred, y_test)
acc
Out[5]:
How about data that do not seem linearly separable?
- We add a non-linear feature makes SVMs capable of linearly separating the data
- As you can see here, it seems that we cannot draw a linear line to split the data
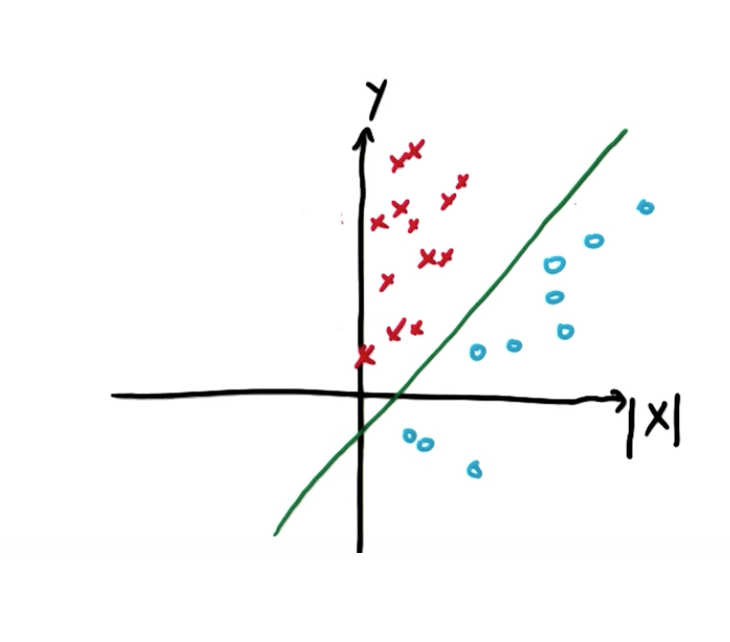
- However, we can add a new feature |x| to split the data
- The new points will be linearly separable
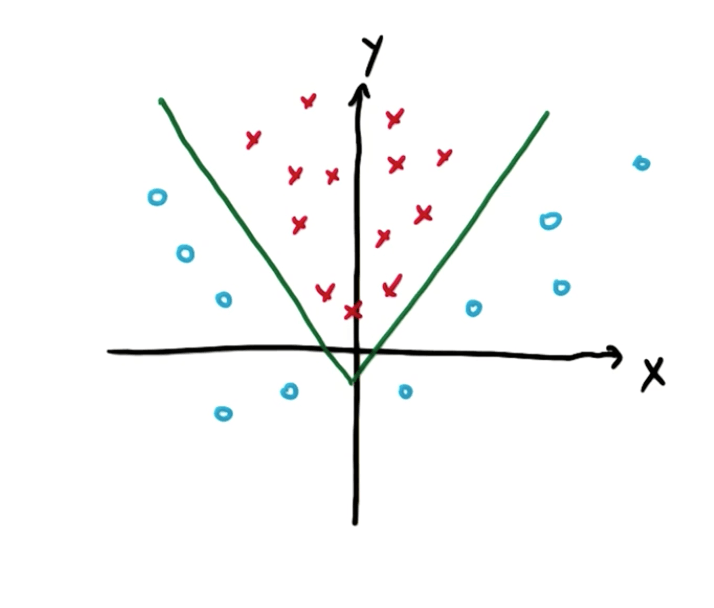
- On the original plot, we can see this is how the data is separated
- As you can see here, it seems that we cannot draw a linear line to split the data
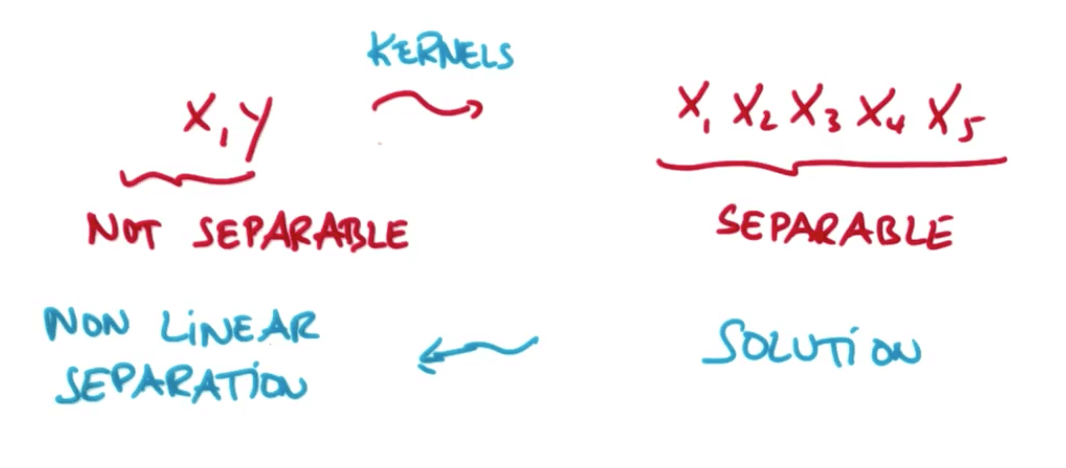
- It seems like we need to create new features, but there's a cool trick called the "kernel trick"
Kernel Trick
- We can map non-separable data to a higher dimensionality to make it separable and then map back
Hyperparameters Tuning with Scikit-Learn
- Kernels
- You can use common kernels and custom kernels
- Common kernels
- 'linear', 'poly', 'rbf', 'sigmoid', 'precomputed'
- Common kernels
- You can use common kernels and custom kernels
- C
- Controls tradeoff between smooth decision boundary and classifying training points correctly
- The C parameter tells the SVM optimization how much you want to avoid misclassifying each training example
- Large C
- The optimizer will choose a smaller-margin hyperplane if that hyperplane does a better job of getting all the training points classified correctly
- Small C
- The optimizer will look for a larger-margin separating hyperplane, even if that hyperplane misclassifies more points
- For very tiny values of C, you should get misclassified examples, often even if your training data is linearly separable
- The optimizer will look for a larger-margin separating hyperplane, even if that hyperplane misclassifies more points
- Large C
- gamma
- Defines how far the influence of a single example reaches
- Low values
- Far
- Only consider close points
- High values
- Low
- Even far-away points get considered
- You can end up with a wiggly decision boundary (over-fitting)
- Low values
- Kernel coefficient for ‘rbf’, ‘poly’ and ‘sigmoid’. If gamma is ‘auto’ then 1/n_features will be used instead.
- Defines how far the influence of a single example reaches
SVMs Suitability
- Does not work well with large datasets due to speed issues
- Does not work well with a lot of noise
- Naive Bayes would be better
- Works well for data that can be linearly classified